Wikipedia
- Details
- Last Updated: Thursday, 22 August 2013 15:18
 General Purpose Coarse-Grained Force Field
General Purpose Coarse-Grained Force Field 

























































































































































|
D.H. de Jong, G. Singh, W.F.D. Bennett, C. Arnarez, T.A. Wassenaar, L.V. Schäfer, X. Periole, D.P. Tieleman, S.J. Marrink
Improved Parameters For The Martini Coarse-Grained Protein Force Field, J. Chem. Th. Comp, 9:687–697, 2013. DOI:10.1021/ct300646g
|
|
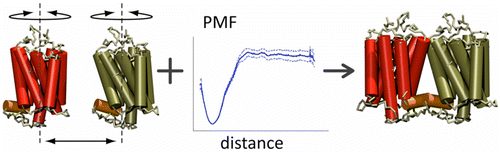
First report on the free energy surface of membrane protein dimerization
A new script to generate both structure and topology files for coarse grain proteins has been released: Martinize.py. Generating input files for a Martini simulation from an atomistic structure takes only one step!
Dimerization of Martini amino acids comparable to atomistic force fields
Martini in Nature